High-Resolution In Vivo Identification of miRNA Targets by Halo-Enhanced Ago2 Pull-Down - ScienceDirect
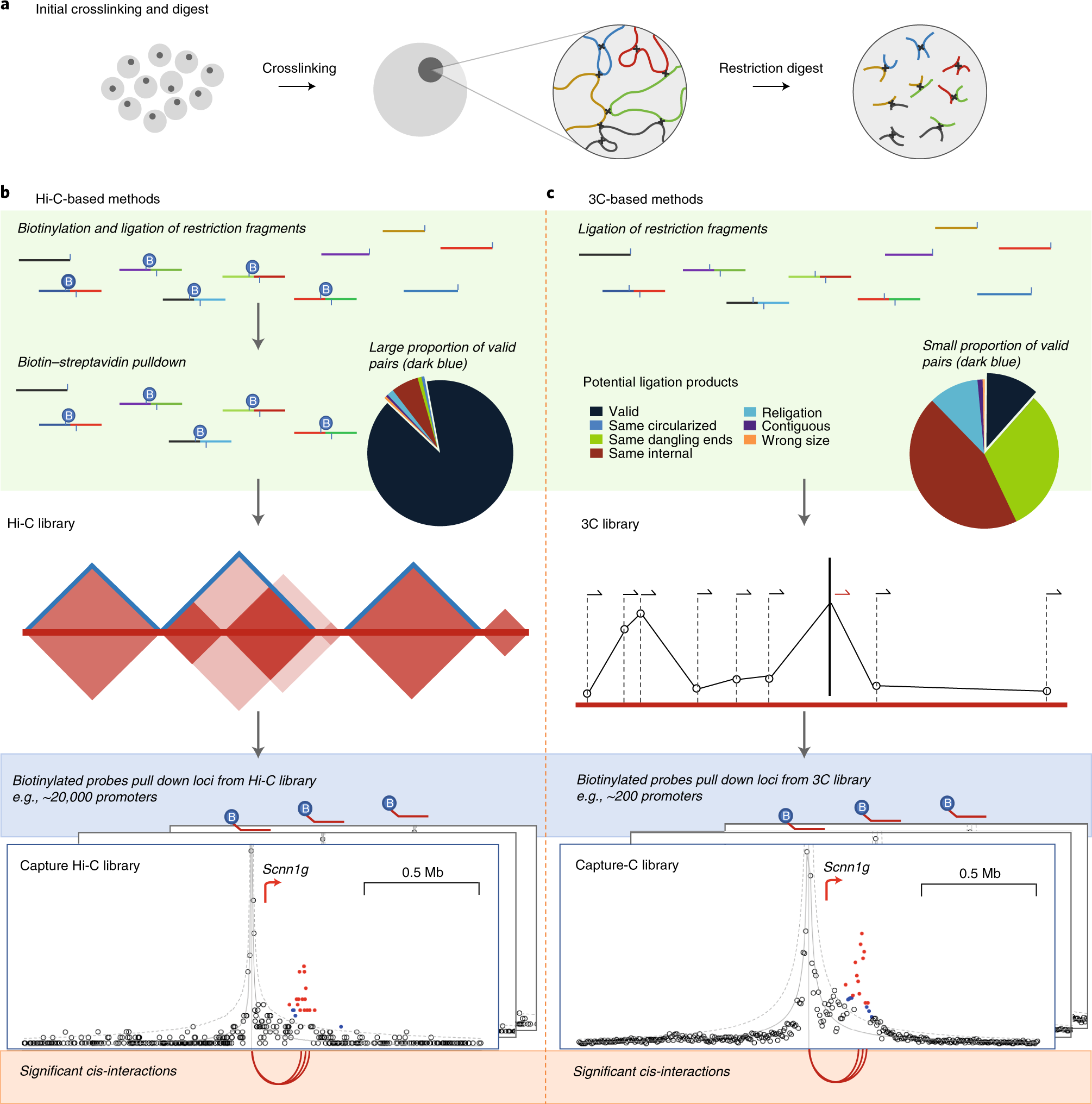
Detecting chromosomal interactions in Capture Hi-C data with CHiCAGO and companion tools | Nature Protocols

Representative multiplex images for all 10 loci in samples from four... | Download Scientific Diagram

SOLVED: Mark the wrong statement about crossing overs (XO): There are 1 to 2 XOs per chromosome pair The XOs help positioning paired chromosomes to the metaphase plate by resisting the pull